If you’ve ever wanted a deeper understanding of what’s going on behind the scenes of correspondence analysis, then this post is for you. Correspondence analysis is a popular tool for visualizing the patterns in large tables. For many practitioners, it is probably a black box. Table goes in, chart comes out. In this post, I explain the mathematics of correspondence analysis. I show each step of the calculation, and I illustrate all the of the steps using R.
If you want to quickly make your own correspondence analysis, this is probably the wrong post for you – but you can easily do that using this template!
Correspondence Analysis in R: A case study
The data that I analyze shows the relationship between thoroughness of newspaper readership by education level. It is a contingency table, which is to say that each number in the table represents the number of people in each pair of categories. For example, the cell in the top-left corner tells us that 5 people with some primary education glanced at the newspaper. The table shows the data for a sample of 312 people (which is also the sum of the numbers displayed).
I show the R code for generating this table below. I have named the resulting table N.
N = matrix(c(5, 18, 19, 12, 3, 7, 46, 29, 40, 7, 2, 20, 39, 49, 16),
nrow = 5,
dimnames = list(
"Level of education" = c("Some primary", "Primary completed", "Some secondary", "Secondary completed", "Some tertiary"),
"Category of readership" = c("Glance", "Fairly thorough", "Very thorough")))
Computing the observed proportions (P) in R
The first step in correspondence analysis is to sum up all the values in the table. I’ve called this total n.
n = sum(N)
Then, we compute the table of proportions, P. It is typical to use this same formula in other types of tables, even if the resulting numbers are not strictly-speaking proportions. Examples include correspondence analysis of tables of means or multiple response data.
P = N / n
This gives us the following table. To make it easy to read, I have done all the calculations in Displayr, which automatically formats R tables using HTML. If you do the calculations in normal R, you will instead get text-based table like the one above.
Row and column masses
In the language of correspondence analysis, the sums of the rows and columns of the table of proportions are called masses. These are the inputs to lots of different calculations. The column masses in this example show that Glance, Fairly thorough, and Very thorough describe the reading habits of 18.3%, 41.3%, and 40.4% of the sample respectively. We can compute the column masses using the following R code:
column.masses = colSums(P)
The row masses are Some primary (4.5%), Primary completed (26.9%), Some secondary (27.9%), Secondary completed (32.4%), and Some tertiary (8.3%). These are computed using:
row.masses = rowSums(P)
Expected proportions (E)
Referring back to the original table of proportions, 1.6% of people glanced and had some primary education. Is this number big or small? We can compute the value that we would expect to see under the assumption that there is no relationship between education and readership. The proportion that glances at a newspaper is 18.2% and 4.5% have only Some primary education. Thus, if there is no relationship between education and readership, we would expect that 4.5% of 18.2% of people (i.e., 0.008 = 0.8%) have both glanced and have primary education. We can compute the expected proportions of all the cells in the table in the same way.
The following R code computes all the values in a single line of code, where %o% means that a table is created by multiplying each of the row totals (row masses) by each of the column totals.
E = row.masses %o% column.masses
Residuals (R)
We compute the residuals by subtracting the expected proportions from the observed proportions. Residuals in correspondence analysis have a different role to that which is typical in statistics. Typically in statistics, the residuals quantify the extent of error in a model. In correspondence analysis, by contrast, the whole focus is on examining the residuals.
The residuals quantify the difference between the observed data and the data we would expect under the assumption that there is no relationship between the row and column categories of the table (i.e., education and readership, in our example).
R = P - E
The biggest residual is -0.045 for Primary completed and Very thorough. That is, the observed proportion of people that only completed primary school and are very thorough is 6.4%, and this is 4.5% lower than the expected proportion of 10.9%, which is computed under the assumption of no relationship between newspaper readership and education. Thus, the tentative conclusion that we can draw from this is that there is a negative association between having completed primary education and reading very thoroughly. That is, people with only primary school education are less likely to read very thoroughly than the average person.
Indexed residuals (I)
Take a look at the top row of the residuals shown in the table above. All of the numbers are close to 0. The obvious explanation for this – that having some primary education is unrelated to reading behavior – is not correct. The real explanation is all the observed proportions (P) and the expected proportions (E) are small because only 4.6% of the sample had this level of education. This highlights a problem with looking at residuals from a table. By ignoring the number of people in each of the rows and columns, we end up being most likely to find results only in rows and columns with larger totals (masses). We can solve this problem by dividing the residuals by the expected values, which gives us a table of indexed residuals (I).
I = R / E
The indexed residuals have a straightforward interpretation. The further the value from the table, the larger the observed proportion relative to the expected proportion. We can now see a clear pattern. The biggest value on the table is the .95 for Some primary and Glance. This tells us that people with some primary education are almost twice as likely to Glance at a newspaper as we would expect if there were no relationship between education and reading. In other words, the observed value is 95% higher than the expected value. Reading along this first row, we see that there is a weaker, but positive, indexed residual of 0.21 for Fairly thorough and Some primary. This tells us that people with some primary education were 21% more likely to be fairly thorough readers that we would expect. And, a score of -.65 for Very thorough, tells us that people with Some primary education were 65% less likely to be Very thorough readers than expected. Reading through all the numbers on the table, the overall pattern is that higher levels of education equate to a more thorough readership.
As we will see later, correspondence analysis is a technique designed for visualizing these indexed values.
Reconstituting indexed residuals from a map
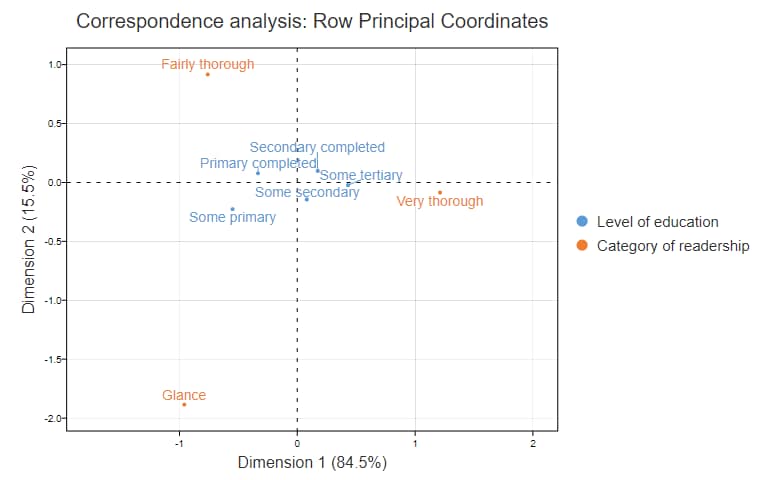
The chart below is a correspondence analysis with the coordinates computed using row principal normalization. I will explain its computation later. Now, I am going to show how we can work backward from this map to the indexed residuals, in much the same way that we can recreate orange juice from orange juice concentrate. Some Primary has coordinates of (-.55, -.23) and Glance’s coordinates are (-.96, -1.89). We can compute the indexed value by multiplying together the two x coordinates and the two y coordinates and summing them up. Thus we have -.55*-.96 + -.23 * -1.89 = .53 + .44 = .97. Taking rounding errors into account, this is identical to the value of .95 shown in the table above.
Unless you have studied some linear algebra, there is a good chance that this calculation, known as the dot product (or a scalar product or inner product), is not intuitive. Fortunately, it can be computed it in a different way that makes it more intuitive.
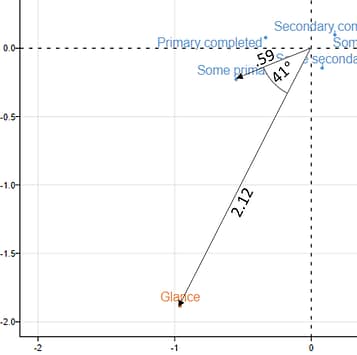
To compute the indexed residual for a couple of points, we start by measuring the distance between each of the points and the origin (see the image to the right). In the case of Some primary, the distance is .59. Then, we compute the distance for Glance, which is 2.12. Then we compute the angle formed when we draw lines from each of the points to the origin. This is 41 degrees. Lastly, we multiply together each of these distances with the cosine of the angle. This gives us .59*2.12*cos(41°) = .59*2.12*.76 = .94. Once rounding errors are taken into account, is the same as the correct value of .95.
Now, perhaps this new formula looks no simpler than the dot product, but if you look at it a bit closer, it becomes pretty straightforward. The first two parts of the formula are the distance of each point from the origin (i.e., the (0,0) coordinate). Thus, all else being equal, the further the point is from the origin, the stronger the associations between that point and the other points on the map. So, looking at the top, we can see that the column category of Glance is the one which is most discriminating in terms of the readership categories.
The second part to interpretation, which will likely bring you back to high school, is the meaning of the cosine. If two points are in exactly the same direction from the origin (i.e., they are on the same line), the cosine of the angle is 1. The bigger the angle, the smaller the cosine, until we get to a right-angle (90° or 270°), at which point the cosine is 0. And, when the lines are going in exactly opposite directions (i.e., so the line between the two points goes through the origin), the cosine of the angle is -1. So, when you have a small angle from the lines connecting the points to the origin, the association is relatively strong (i.e., a positive indexed residual). When there is a right angle there is no association (i.e., no residual). When there is a wide angle, a negative residual is an outcome.
Putting all this together allows us to work out the following things from the row principal correspondence analysis map above, which I have reproduced below to limit scrolling:
- People with only Primary completed are relatively unlikely to be Very thorough.
- Those with Some primary are more likely to Glance.
- People with Primary completed are more likely to be Fairly thorough.
- The more education somebody has, the more likely they are to be Very thorough.
Reconstituting residuals from bigger tables
If you look at the chart above, you can see that it shows percentages in the x and y labels. (I will describe how these are computed below.) They indicate how much of the variation in the indexed residuals is explained by the horizontal and vertical coordinates. As these add up to 100%, we can perfectly reconstitute the indexed residuals from the data. For most tables, however, they add up to less than 100%. This means that there is some degree of information missing from the map. This is not unlike reconstituted orange juice, which falls short of fresh orange juice.
The post How to Interpret Correspondence Analysis Plots (It Probably Isn’t the Way You Think) provides a much more thorough (but un-mathematical) description of issues arising with the interpretation of correspondence analysis.
Singular values, eigenvalues, and variance explained
In the previous two sections, I described the relationship between the coordinates on the map and the indexed residuals. In this section, I am going to explain how the coordinates are computed from the indexed residuals.
The first step in computing the coordinates is to do a near-magical bit of mathematics called a Singular Value Decomposition (SVD). I have had a go at expressing this in layperson’s language in my post An Intuitive Explanation of the Singular Value Decomposition (SVD): A Tutorial in R, which works through the same example that I have used in this post.
The code that I used for performing the SVD of the indexed residuals is shown below. The first line computes Z, by multiplying together each of indexed residuals by the square root of their corresponding expected values. This seems a bit mysterious at first, but two interesting things are going on here.
First, Z is a standardized residual, which is a rather cool type of statistic in its own right. Second, and more importantly from the perspective of correspondence analysis, what this does is cause the singular value decomposition to be weighted, such that cells with a higher expected value are given a higher weight in the data. As often the expected values are related to the sample size, this weighting means that smaller cells on the table, for which the sampling error will be larger, are down-weighted. In other words, this weighting makes correspondence analysis relatively robust to outliers caused by sampling error, when the table being analyzed is a contingency table.
Z = I * sqrt(E) SVD = svd(Z) rownames(SVD$u) = rownames(P) rownames(SVD$v) = colnames(P)
A singular value decomposition has three outputs:
- A vector, d, contains the singular values.
- A matrix u which contains the left singular vectors.
- A matrix v with the right singular vectors.
The left singular vectors correspond to the categories in the rows of the table and the right singular vectors correspond to the columns. Each of the singular values, and the corresponding vectors (i.e., columns of u and v), correspond to a dimension. As we will see, the coordinates used to plot row and column categories are derived from the first two dimensions.
Squared singular values are known as eigenvalues. The eigenvalues in our example are .0704, .0129, and .0000.
eigenvalues = SVD$d^2
Each of these eigenvalues is proportional to the amount of variance explained by the columns. By summing them up and expressing them as a proportion, which is done by the R function prop.table(eigenvalues), we compute that the first dimension of our correspondence analysis explains 84.5% of the variance in the data and the second 15.5%, which are the numbers shown in x and y labels of the scatter plot shown earlier. The third dimension explains 0.0% of the variance, so we can ignore it entirely. This is why we are able to perfectly reconstitute the indexed residuals from the correspondence analysis plot.
Standard coordinates
As mentioned, we have weighted the indexed residuals prior to performing the SVD. So, in order to get coordinates that represent the indexed residuals we now need to unweight the SVD’s outputs. We do this by dividing each row of the left singular vectors by the square root of the row masses (defined near the beginning of this post):
standard.coordinates.rows = sweep(SVD$u, 1, sqrt(row.masses), "/")
This gives is the standard coordinates of the rows:
We do the same process for the right singular vectors, except we use the column masses:
standard.coordinates.columns = sweep(SVD$v, 1, sqrt(column.masses), "/")
This gives us the standard coordinates of the columns, shown below. These are the coordinates that have been used to plot the column categories on the maps we in this post.
Principal coordinates
The principal coordinates are the standard coordinates multiplied by the corresponding singular values:
principal.coordinates.rows = sweep(standard.coordinates.rows, 2, SVD$d, "*")
The positions of the row categories shown on the earlier plots are these principal coordinates. The principal coordinates for the education levels (rows) are shown in the table below.
The principal coordinates represent the distance between the row profiles of the original table. The row profiles are shown in the table below. They are the raw data (N) divided by the row totals. Outside of correspondence analysis, they are more commonly referred to as the row percentages of the contingency table. The more similar two rows’ principal coordinates, the more similar their row profiles. More precisely, when we plot the principal coordinates, the distances between the points are chi-square distances. These are the distances between the rows weighted based on the column masses.
The principal coordinates for the columns are computed in the same way:
principal.coordinates.columns = sweep(standard.coordinates.columns, 2, SVD$d, "*")
In the row principal plot shown earlier, the row categories’ positions are the principal coordinates. The column categories are plotted based on the standard coordinates. This means that it is valid to compare row categories based on their proximity to each other. It is also valid to understand the relationship between the row and column coordinates based on their dot products. But, it is not valid to compare the column points based on their position. I discuss this in more detail in a post called Normalization and the Scaling Problem in Correspondence Analysis: A Tutorial Using R.
Quality
We have already looked at one metric of the quality of a correspondence analysis: the proportion of the variance explained. We can also compute the quality of the correspondence analysis for each of the points on a map. Recall that the further a point is from the origin, the greater that point is explained by the correspondence analysis. When we square the principal coordinates and express these as row proportions, we get measures of the quality of each dimension for each point. Sometimes these are referred to as the squared correlations and squared cosines.
pc = rbind(principal.coordinates.rows, principal.coordinates.columns) prop.table(pc ^ 2, 1)
The quality of the map for a particular category is usually defined as the sum of the scores it gets for the two dimensions that are plotted. In our example, these all add up to 100%.
Acknowledgments
The data in the example comes from Greenacre and Hastie’s 1987 paper “The geometric interpretation of correspondence analysis”, published in the Journal of the American Statistical Association.
Where practical, I have used the notation and terminology used in Michael Greenacre’s (2016) third edition of Correspondence Analysis in Practice. This excellent book contains many additional calculations for correspondence analysis diagnostics. The only intentional large deviation from Greenacre’s terminology relates to the description of the normalizations (I discuss the differences in terminology in Normalization and the Scaling Problem in Correspondence Analysis: A Tutorial Using R).
This post is partly based on a paper that I wrote for the International Journal of Market Research, “Improving the display of correspondence analysis using moon plots”, in 2011.